Tag: Self-assembling nanomaterials
-
Designed Protein Containers Push Bioengineering Boundaries
Earlier this month, Baker lab researchers reported the computational design of a hyperstable 60-subunit protein icosahedron in Nature (Hsia et al); icosahedral protein structures are commonly observed in natural biological systems for packaging and transport (e.g. viral capsids). The described design was composed of 60 trimeric protein building blocks that self-assembled in a nanocage. In…
-
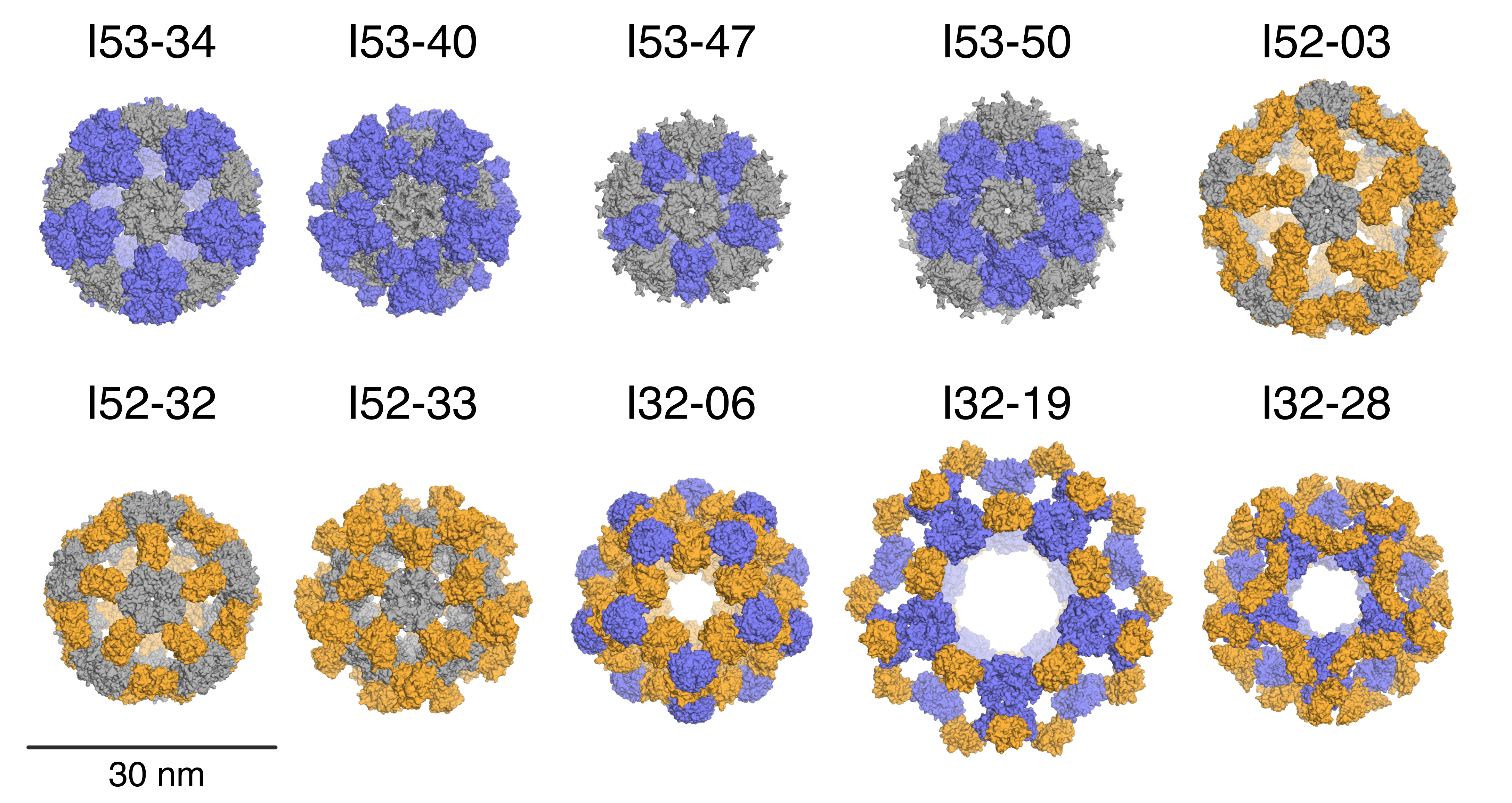
Accurate Design of Co-Assembling Multi-Component Protein Nanomaterials
A new paper is out in the June 5 issue of Nature entitled Accurate design of co-assembling multi-component protein nanomaterials. Scientists at the Institute for Protein Design (IPD), in collaboration with researchers at UCLA and HHMI, have built upon their previous work constructing single-component protein nanocages and can now design and build self-assembling protein nanomaterials made up of…
-
Computational Design of Self-Assembling Protein Nanomaterials with Atomic Level Accuracy
IPD researchers in the Baker group have published in Science a paper entitled “Computational design of self-assembling protein nanomaterials with atomic level accuracy.” They describe a general computational method for designing proteins that self-assemble to a desired symmetric architecture. Protein building blocks are docked together symmetrically to identify complementary packing arrangements, and low-energy protein-protein interfaces are then designed…
