Software
RFdiffusion

Generative AI for protein design
Inspired by AI image generators, RFdiffusion can be used to create novel protein structures in seconds. It sculpts clouds of disconnected atoms into novel protein backbones, yielding monomers, oligomers, binders, and more.
Initially trained only on amino acids, RFdiffusion All-Atom now builds molecules using all of life’s building blocks, including DNA, RNA, ions, and small molecules.
Krishna R, Wang Y, Ahern W, et al. Generalized biomolecular modeling and design with RoseTTAFold All-Atom. Science, 2024
Watson JL, Juergens D, Bennett NR, Trippe BL, Yim J, Eisenach HE, Ahern W, et al. De novo design of protein structure and function with RFdiffusion. Nature, 2023
ProteinMPNN

Rapid sequence design
ProteinMPNN takes a protein structure as input and quickly creates new amino acid sequences that are likely to fold into that backbone. Combined with structure generation tools like RFdiffusion, it can be used to design proteins with never-before-seen sequences, structures, and functions.
ProteinMPNN requires no expert customization, runs in about one second, and outperforms our prior best tools in laboratory tests.
Dauparas J, et al. Atomic context-conditioned protein sequence design using LigandMPNN. Nature Methods, 2025
Dauparas J, et al. Robust deep learning–based protein sequence design using ProteinMPNN. Science, 2022
RoseTTAFold

Protein structure prediction
RoseTTAFold uses multiple neural networks to quickly and accurately predict protein structures based on amino acid sequences. Without the aid of such software, it can take years of laboratory work to measure the shape of just one protein. With RoseTTAFold, a structure can be computed in minutes.
But proteins rarely act alone. That’s why we created RoseTTAFold All-Atom to model all of life’s building blocks. We’ve shown that it can predict in detail how proteins interact with particular DNA stretches, how drug molecules may bind to human receptors, and more.
Krishna R, Wang Y, Ahern W, et al. Generalized biomolecular modeling and design with RoseTTAFold All-Atom. Science, 2024
Baek M, et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science, 2021
Foldit
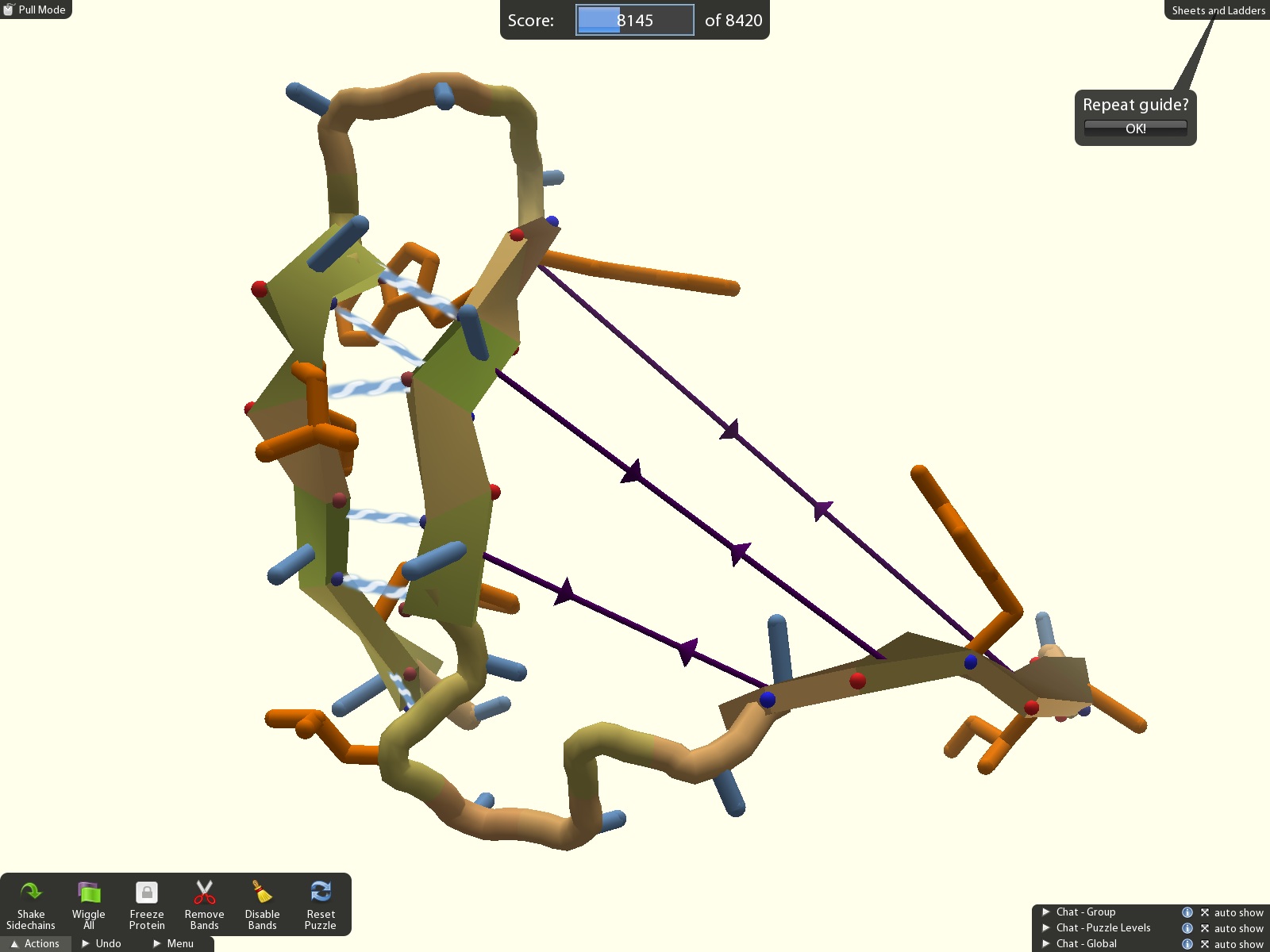
Our free video game
Foldit is a one-of-a-kind computer game developed by university scientists. Every week, the Foldit team post new puzzles focused on the latest problems in protein science, including protein design to treat diseases like influenza and COVID-19, small molecule design to invent new drug compounds, and protein structure solving to map the molecules that drive biology.
Foldit is free to play and not-for-profit. Discoveries made in the game are published in peer-reviewed research journals and Foldit players are always credited for their contributions.
Koepnick B, et al. De novo protein design by citizen scientists. Nature, 2019
Online game Foldit helps anti-AIDS drug quest. BBC, 2011
Cooper S, et al. Predicting protein structures with a multiplayer online game. Nature, 2010
Visit our blog to discover our most recent work.
