In the early 1990s, researchers in the field of protein structure prediction were challenged by the problem of how to impartially judge the accuracy of prediction algorithms. This realization led the protein structure prediction the community to start the Critical Assessment of protein Structure Prediction (CASP), a community-wide, worldwide experiment for protein structure prediction taking place every two years since 1994. In each CASP, an independent scientific advisory board solicits other researchers to submit experimentally verified, but unpublished, 3D protein structures to CASP. The linear amino acid sequences of these proteins are then provided to structure prediction researchers, who each have an equal and limited amount of time to submit final structure predictions to the CASP advisory board. The submitted structure predictions are then compared to the experimentally verified structures using the same metrics for all CASP contributors. Even though the primary goal of CASP is to help advance methods for identifying protein 3D structure given only its linear amino acid sequence, many view the experiment more as a “world championship” in protein structure prediction.
In the early 1990s, researchers in the field of protein structure prediction were challenged by the problem of how to impartially judge the accuracy of prediction algorithms. This realization led the protein structure prediction the community to start the Critical Assessment of protein Structure Prediction (CASP), a community-wide, worldwide experiment for protein structure prediction taking place every two years since 1994. In each CASP, an independent scientific advisory board solicits other researchers to submit experimentally verified, but unpublished, 3D protein structures to CASP. The linear amino acid sequences of these proteins are then provided to structure prediction researchers, who each have an equal and limited amount of time to submit final structure predictions to the CASP advisory board. The submitted structure predictions are then compared to the experimentally verified structures using the same metrics for all CASP contributors. Even though the primary goal of CASP is to help advance methods for identifying protein 3D structure given only its linear amino acid sequence, many view the experiment more as a “world championship” in protein structure prediction.
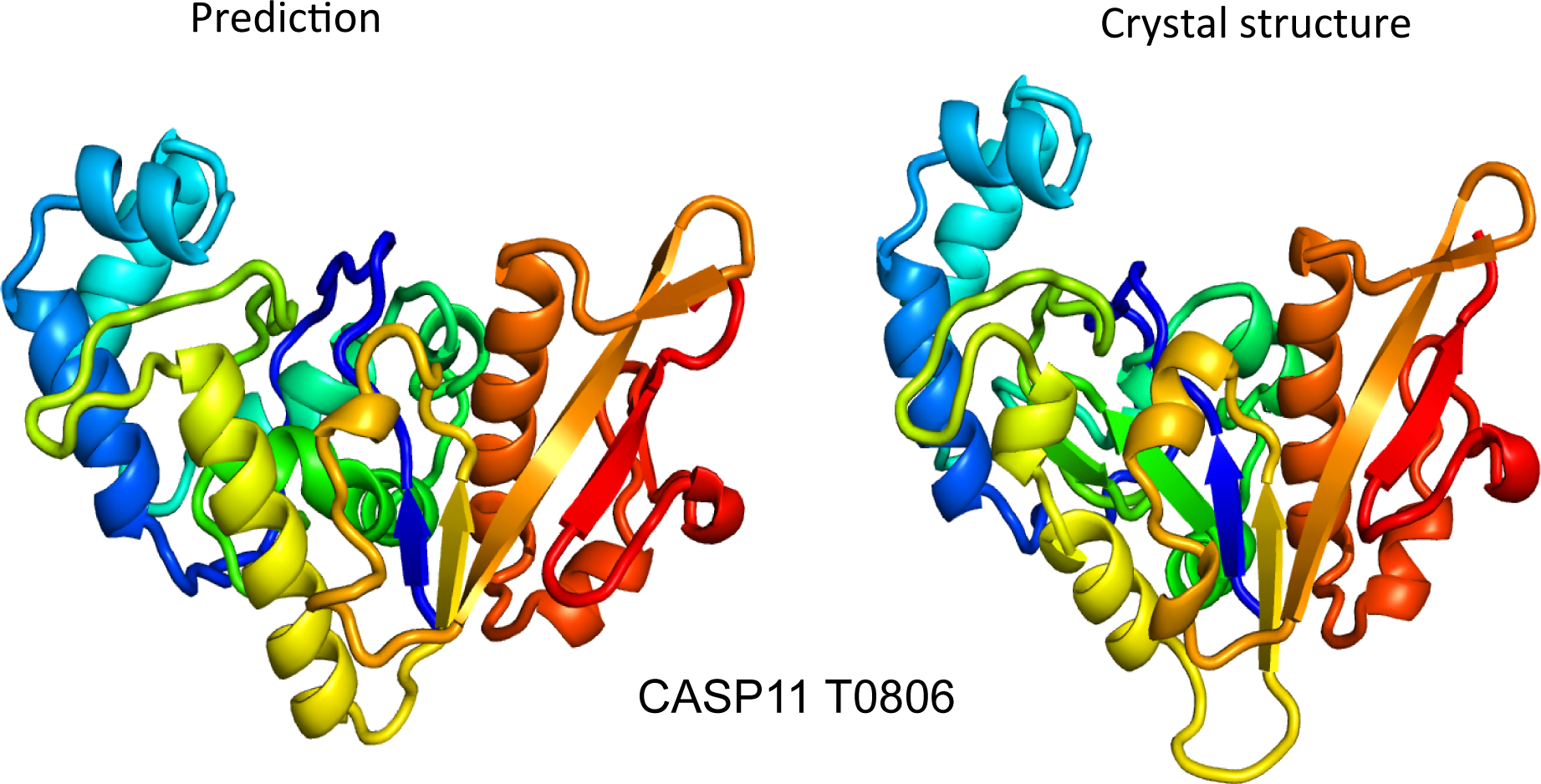
Over a 16 year period (CASP3-11), the Baker lab has consistently achieved top performance in the hardest category of structure prediction; the “Twilight Zone” where the linear amino acid sequence of the protein shares no discernable relation to any publicly available 3D structure. In 2014 this culminated in our highly accurate blind structure predictions of two large proteins each >200 amino acids in length. Our methods involve using DNA sequence information to help us predict the 3D structures of proteins.
We recently published these results in E-life, and the results are getting significant attention.
You can read the general-public summary here:
http://elifesciences.org/content/4/e09248/abstract-2
Or find the whole thing here:
http://elifesciences.org/content/4/e09248
Also, learn about the first author of the paper, Sergey Ovchinnikov, by watching this interview: