In the summer of 1961, Osamu Shimomura drove across the country in a cramped station wagon to scoop jellyfish from the docks of Friday Harbor. He wanted to discover what made them glow.
It took Shimomura and other biochemists more 30 years to find a full answer. By then, recombinant DNA technology allowed researchers to clone and characterize the two proteins responsible: aequorin and GFP. The latter would earn Shimomura his share of the 2008 Nobel Prize.
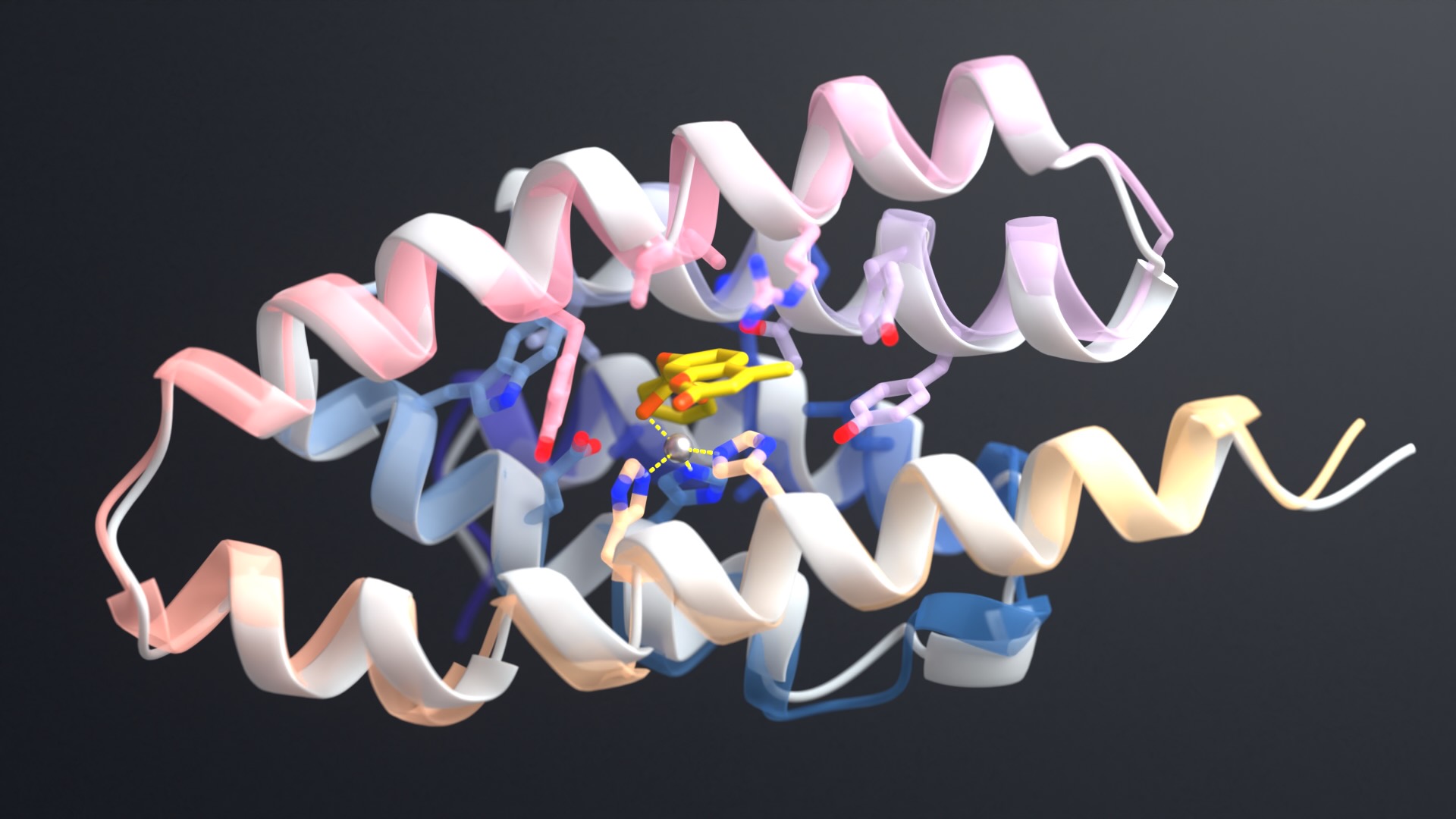
GFP, a 238-residue beta-barrel with a covalently linked chromophore, transformed how scientists study cells and the molecules in them. As a genetic tag, GFP has illuminated the inner workings of human brain cells, bacteria, fungi and more.
This week, scientists from the IPD report in Nature the design of a completely artificial fluorescent beta-barrel protein.
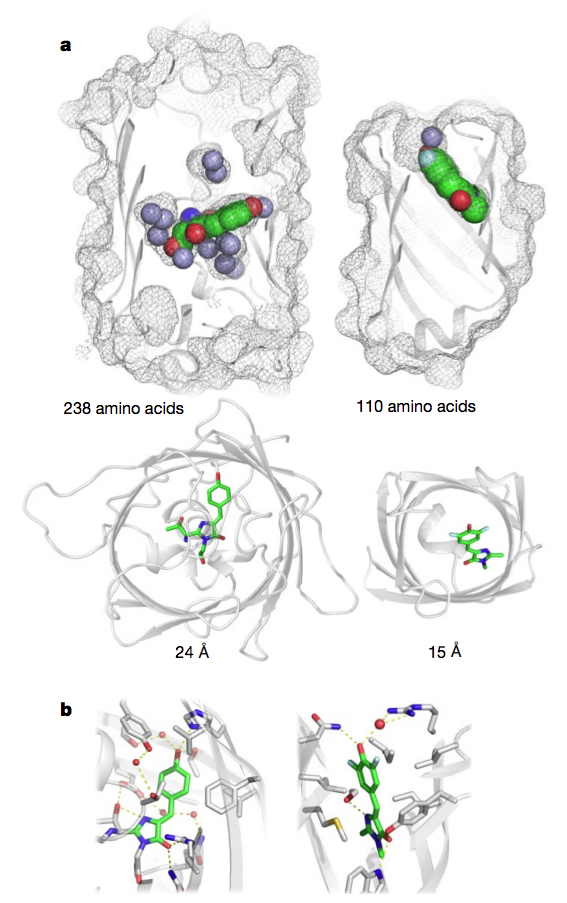
mesh and ribbon representations. b, Close-up of chromophore
binding interactions.
Many natural proteins evolved to bind small molecules. Reengineering such proteins is rarely straightforward, limiting how they can be applied. The new findings demonstrate that proteins unlike any found in nature can be rationally-designed to bind to and act on specific small molecules with high precision and affinity.
The lead authors of the paper are Jiayi Dou, Ph.D and Anastassia A. Vorobieva, Ph.D., then both senior fellows in the Baker lab.

Anastassia Vorobieva with her son Alexandre (left) and Jiayi Dou (right).
To make the fluorescent protein the researchers had to achieve another first: Creating beta-barrels from scratch. The fold was ideal because one end of its cylindrical shape could be used to stabilize the protein while the other could be used to create a cavity that would serve as the binding site for the target molecule, DFHBI. In nature, beta-barrels proteins bind a wide range of small molecules.
Rosetta was used to design the scaffold de novo. To create the cavity, the team used a new docking algorithm called the “Rotamer Interaction Field” or RIF, developed by William Sheffler, Ph.D., a senior research scientist in the Baker lab, that rapidly identifies all potential structures of cavities that fulfill the prerequisites for binding specific molecules.
The designed protein absorbs blue and emits cyan light. It is stable up to 75°C.
“It worked in bacterial, yeast and mammalian cells,” said Dou, “and being half the size of green fluorescent protein should be very useful to researchers.”
“Equally important,” Baker added, “it greatly advances our understanding of the determinants of protein folding and binding beyond what we have learned from describing existing protein structures.”
Read the article here: https://www.nature.com/articles/s41586-018-0509-0
View the PDF here: Fluorescent proteins designed from scratch