This week we report in JACS a general approach for designing self-assembling 2D protein arrays. This project was led by Zibo Chen, a recent Baker lab graduate student, and featured collaborators from the, DiMaio, De Yoreo and Kollman labs at UW.
Abstract:
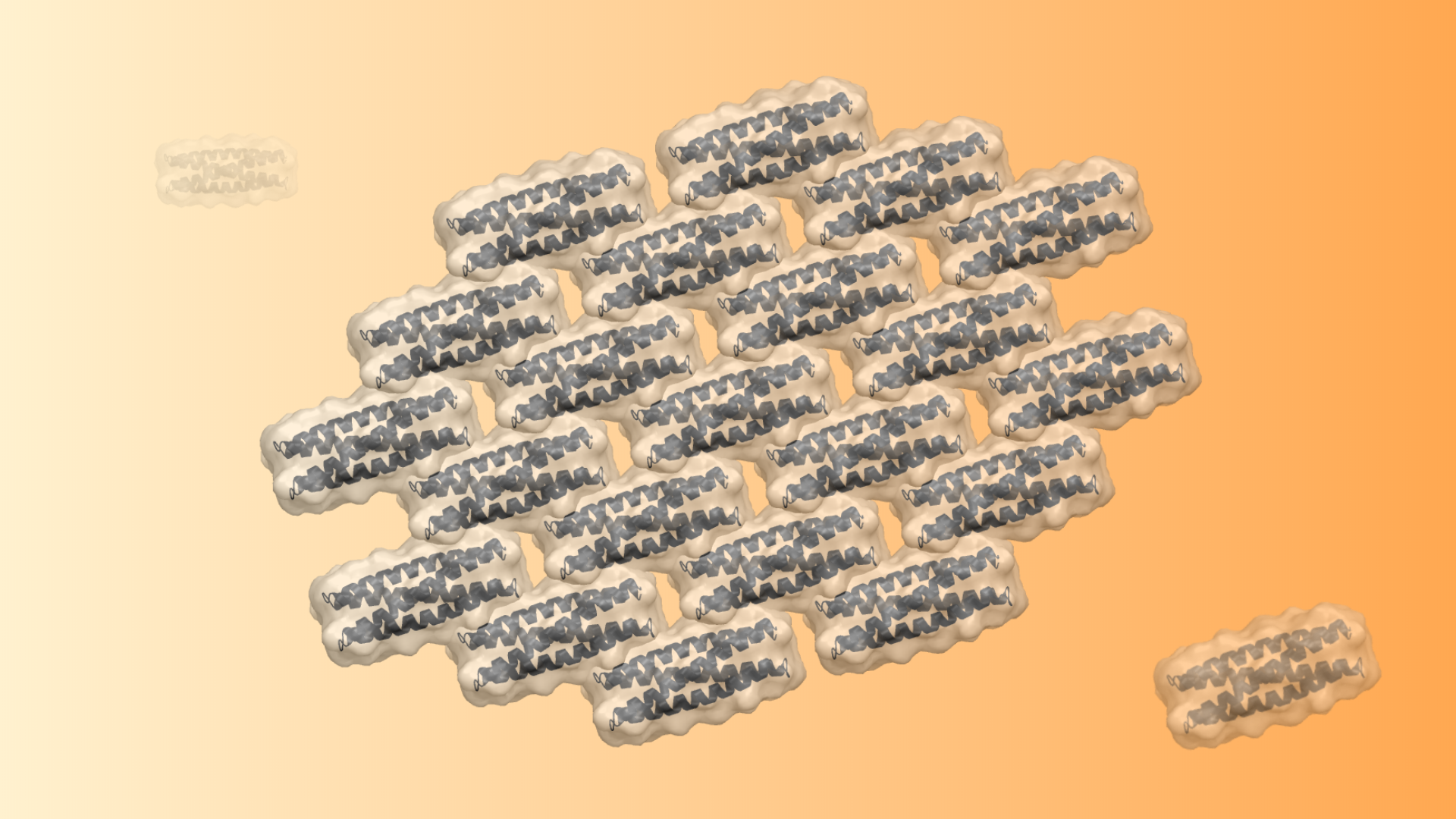
Modular self-assembly of biomolecules in two dimensions (2D) is straightforward with DNA but has been difficult to realize with proteins, due to the lack of modular specificity similar to Watson-Crick base pairing. Here we describe a general approach to design 2D arrays using de novo designed pseudosymmetric protein building blocks. A homodimeric helical bundle was reconnected into a monomeric building block, and the surface was redesigned in Rosetta to enable self-assembly into a 2D array in the C 1 2 layer symmetry group. Two out of ten designed arrays assembled to μm scale under negative stain electron microscopy, and displayed the designed lattice geometry with assembly size up to 100 nm under atomic force microscopy. The design of 2D arrays with pseudosymmetric building blocks is an important step toward the design of programmable protein self-assembly via pseudosymmetric patterning of orthogonal binding interfaces.
Read the full report here: https://pubs.acs.org/doi/abs/10.1021/jacs.9b01978# PDF