The latest paper coming out from the IPD was published today on the Science website. It’s titled “Principles for designing proteins with cavities formed by curved β sheets” with first co-authors Enrique Marcos and Benjamin Basanta, a former and current IPD member, respectively. Other IPD members on the paper include Tamuka Chidyausiku, Gustav Oberdorfer, Daniel-Adriano Silva, Jiayi Dou, and David Baker. Dr. Baker wrote a summary about the publication:
Some of the key functions of the proteins in our bodies and in all living things are to catalyze chemical reactions—speed up the rates by many orders of magnitude-and to sense and respond to small molecules in the body and in the environment. New proteins that catalyze chemical reactions and/or sense and respond to compounds not found in nature would have wide use in medicine and industry.

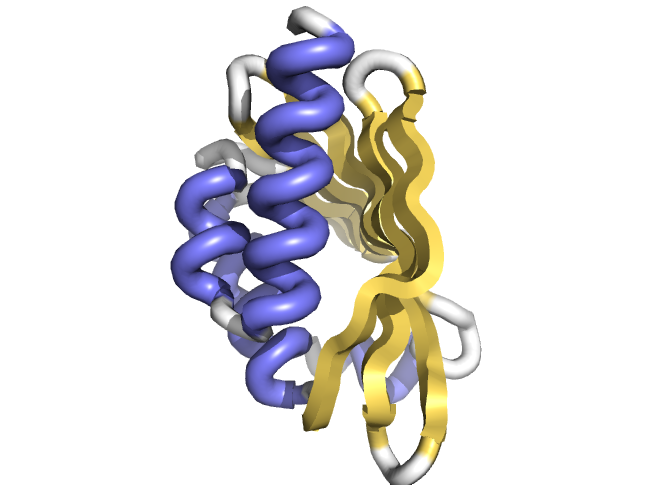
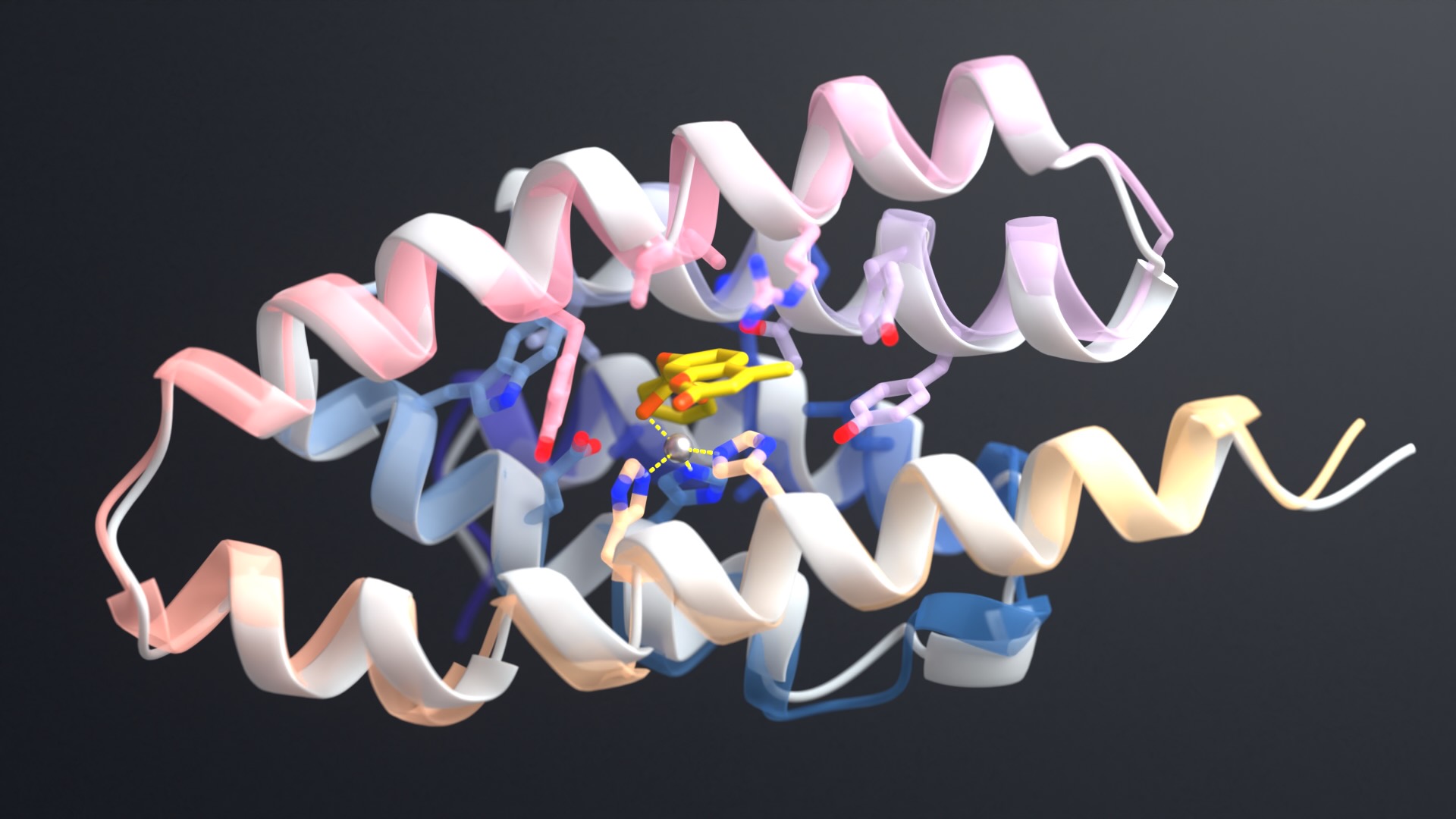
Computational protein design can in principle be used to generate such new catalysts and receptors, but a major challenge to accomplishing this has been the inability to design proteins with cavities within which the catalysis or small molecule binding can take place. This paper describes a general approach for designing proteins with cavities with tunable size and shape. The method opens the door to design of new catalysts and binding proteins [by generating proteins with appropriately sized and shaped cavities to hold the small molecule and lining the cavity with amino acid functional groups to carry out catalysis and/or binding].
Read the UW’s HS NewsBeat write-up here.